In a three-piece blog series, we introduce you to our FAIR Data Station. In this first part, we explain how to get started with Data FAIRification, the easy way.
By the FAIR Data Platform / November 16, 2021
KEY MESSAGES
- The FAIR guiding principles encourage scientists to find, access, interoperate, combine and reuse data in an easy way.
- In general, with data comes metadata, which is best to add and update right from the start
- Our FAIR Data Station guides researchers in metadata management via a simple three-step process, as we describe in this blog.
- You can also access the FAIR Data Station directly here
The ‘FAIR Guiding Principles for scientific data management and stewardship’ are built upon the use of machine-actionable metadata to find, access, interoperate, combine, and reuse data with minimal human intervention. The amount and level of detail of this metadata must be sufficient to allow for unambiguous interpretation of the associated data. To maximize the potential for reuse, the Life Sciences use Minimum Information (MI) standards, which consist of two parts, namely:
- First, each sample/assay and its associated data type comes with a community-accepted checklist of (mandatory) reporting requirements.
- Next, to ensure metadata machine-actionability, such data should be reported in a community-accepted data format. For example, dates should be supplied in the ISO8601 format (YYYY-MM-DD)
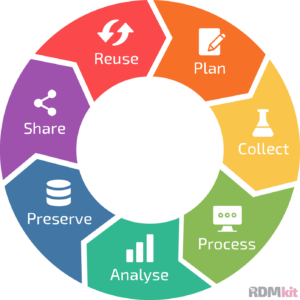

The data life cycle
In UNLOCK, data management goes hand in hand with metadata management. This implies that we work according to the FAIR-by-design principles, in which project metadata is added and continuously updated from the start. This is not only crucial to obtain reusable data objects, but also because we believe that adding metadata from the start helps to increase the quality and the reproducibility of research. As seen in the data life cycle (Figure 1), several steps, such as experiment planning, sample collection, processing, and analysis typically require human involvement. It is therefore essential that the metadata associated with these steps can easily be captured and integrated with the research workflows. To this end, we have developed the FAIR Data Station, a metadata ingestion platform that helps the researcher to improve the quality and safeguards the machine-actionability of experiment metadata. In brief, data FAIRification the easy way!
Data FAIRification: A three-step process
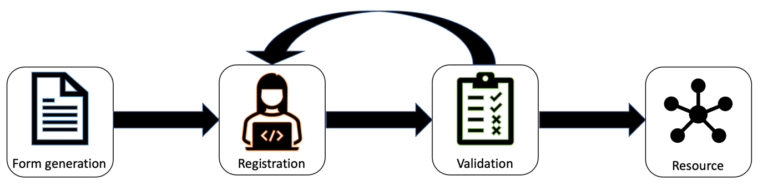
The FAIR Data Station guides researchers in metadata management in a simple three-step process, as depicted in Figure 2.
1. Firstly, selection of the appropriate metadata standard(s), which results in the generation of a template spreadsheet in standard Excel format with a machine-actionable header.
- This Excel sheet is used for registration of sample and assay metadata, which is done offline.
2. Secondl, validation of sample and assay metadata content (i.e. completeness and format) according to requirements of the selected MI-standard.
- To this end, the Excel file is uploaded to the FAIR-Data Station. Meanwhile, it is automatically checked for inconsistencies.
3. Lastly, generation of FAIR machine-actionable metadata in the RDF data model.
- Accepted Excel files will automatically be converted in a Resource Description Framework (RDF) data model that can be queried using the SPARQL query language.
Mininum Information standards
The FAIR Data Station can be used with any MI standard, but by default it is loaded with the Minimum Information about any (x) Sequence (MIxS) standard developed by the Genomic Standards Consortium (see this Nature Biotechnology paper for motivations). The latest version of these MIxS standards can be obtained here. As mentioned previously, sample and assay registration is done offline using the preformatted Excel spreadsheet. In fact, this setup offers advantages over web-based methods when dealing with time series, when you have many samples or multiple assays and, additionally, when your experiments involves multiple researchers. Note that the workflow also allows for routine intermediate validation of registered samples and assays.
Accessing the FAIR Data Station
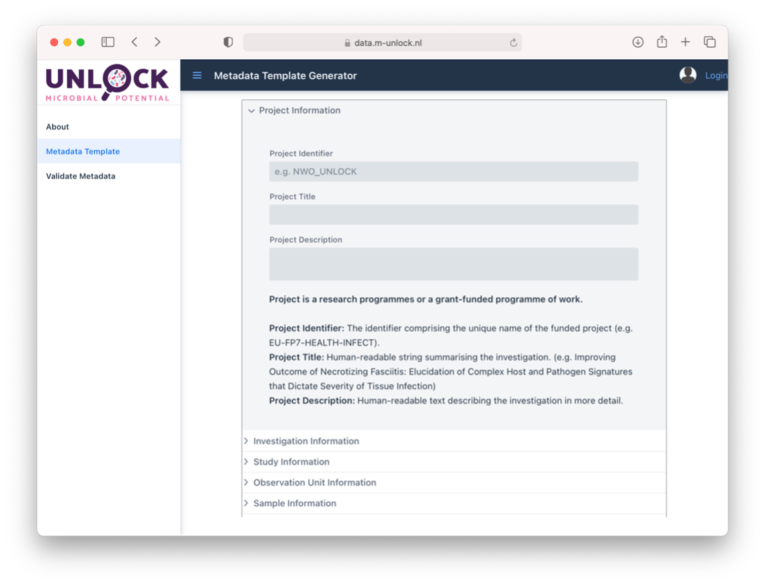
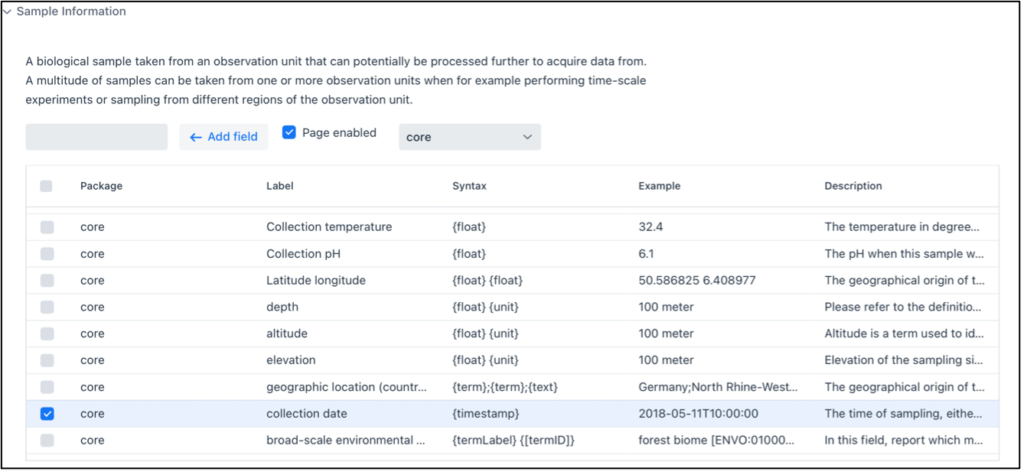
You can access the FAIR Data Station via this link. First, the project context metadata (project information, experimental aim and setup) is collected using the web interface (Figure 3).Next, thhis metadata is converted in the Investigation/Study/Assay (ISA) data model, after which the appropriate MI-standard is selected. A MI-standard often consists of a checklist of mandatory, conditional mandatory, environment-dependent and optional reporting requirements. Mandatory items are selected by default and can thus not be deselected. Here, the researcher selects the relevant optional reporting metadata requirements. After completion of the webform, a preformatted multi-sheet Excel file is generated each representing an ISA level. The Sample and Assay worksheets hold sample details as well as analytical measurement details. They also have have machine-actionable column headers representing the mandatory and user-selected optional elements. Note that for convenience free-text comment columns can be added to these worksheets as well.
Useful links
In case you want to get started, go directly to the FAIR Data Station. Still questions or suggestions? Do not hesistate to contact us.
In the next blog, we further clarify why you should bother about data FAIRification. Finally, we will end this series by explaining how you can build your own Semantic database.
